All published articles of this journal are available on ScienceDirect.
Endoplasmic Reticulum (ER) Stress Enhances Tip60 (A Histone Acetyltransferase) Binding to the Concanavalin A
Abstract
Herein, we report that the concanavalin A binding of Tip60 (a target of the human immunodeficiency virus type 1-encoded transactivator Tat interacting protein 60 KD; a histone acetyltransferase; HAT) is enhanced as the result of endoplasmic reticulum (ER) stress. The cell expression of Tip60 combined with site-directed mutagenesis analysis was used to identify the glutamine 324 residue as the lecithin binding (Concanavalin A; Con A) site. The Tip60 N324A mutant strain, which seems to be the Con A binding-deficient, was attenuated the protein-protein interactions with FE65 and its protein stability, but its ability of G0-G1 cell cycle arrest was not interrupted. Interestingly, both HAT activity and the nuclear localization of Tip60 N324A mutant were enhanced than those of Tip60 WT. Thus, our results indicate that the Con A binding deficient of Tip60 seems to be one of the most pivotal posttranslational modifications (such as N-glycosylation) for its functional regulation signal, which is generated in response to ER stress.
INTRODUCTION
Tip60 was characterized as a partner of the human immunodeficiency virus, type 1-encoded transactivator protein Tat. Tip60’s crucial role in cell signaling is probably the principal reason for its targeting by the HIV-1-encoded transactivator Tat [1, 2]. It appears that Tat employs Tip60 to control cellular events for the benefit of the virus [3]. Tip60 was subsequently identified as a specific member of the MYST family of nuclear histone acetyltransferases (HATs). While attempting to elucidate its function and identify proteins associated with ectopically expressed Tip60, the presence of associated ATPase and DNA helicase activities was demonstrated [4, 5]. Functional tests demonstrated that Tip60 and its associated proteins may perform essential functions in DNA repair and apoptosis. Moreover, Tip60 appears to be involved in a broad variety of cellular functions [6, 7].
N-glycosylation is one of the most common and yet complex forms of post-translational modification in the endoplasmic reticulum (ER) and Golgi body [8, 9]. It is involved intricately in a variety of cellular processes, including protein folding, protein secretion, intracellular trafficking, stability, binding affinity, enzyme activity, and substrate specificity, enabling the fine-tuning of a protein's function [10, 11]. N- glycosylation is a process in which glycan is attached to a protein at an asparagine residue called the N-X-S/T motif; X here denotes any amino acid except proline [12, 13]. Over half of all proteins in nature are glycosylated, and more than three-quarters of those proteins harbor an N-linked glycan [11, 14].
It has been also revealed that an imbalance between the protein-folding load and the capacity of ER is created, resulting in the accumulation of unfolded or misfolded proteins in the ER lumen - the condition referred to ‘ER stress’ [14-16]. With the accumulation of unfolded or misfolded protein in the ER, the cells respond to activate a group of signal transduction pathways collectively termed Unfolded Protein Response (UPR) to alter transcriptional and translational programs [17, 18]. A growing body of evidence suggests that ER stress, oxidative stress, and the inflammatory response are cross-linked and that limiting of either one affect the others [17, 19]. Even though the scope and consequence of ER stress and the UPR have been significantly investigated, it is still unclear how ER stress transducers its signal to its downstream components step by step.
Here, we reported new evidence of Tip60 (Tip60 isoform 1; 546 aa; Accession Number NP-874369) regulation by N-glycosylation on its glutamine 324 residue, which does not match the consensus motif for the asparagine (N) residue of the NX(S/T). First, Tip60 has a property to bind in vivo with Con A (Jack bean lectin). Second, the major influence site of Tip60 to bind Con A has been discovered with site-directed mutagenesis. We demonstrated that the Gln-324 residue of Tip60 also seems to be specifically the N-glycosylated site, and that the Con A binding of Tip60 was blocked with a specific inhibitor of glycosylation (tunicamycin). Third, we determined that the interaction between FE65 and Tip60 occurs through the PID1 of Fe65 and the 324NEIY327 motif of Tip60 which is overlapped with the glycosylation of Tip60. Forth, we noted that Con A binding to the site of Tip60 is related with its protein stability, subcellular localization, and HAT activity. Finally, we observed that ER (endoplasmic reticulum) stress induced by brefeldin A promotes the Con A binding site of Tip60. Thus, our data lead us to the novel suggestion that Con A binding site on the Gln-324 residue of Tip60 (a putative N-glycosylation site), which is involved in the response to ER stress, constitutes one of the major posttranslational modifications for the control of its function.
MATERIALS AND METHODS
Production and Purification of Proteins
Wild-type or mutant Tip60 cDNA was cloned into the pCMV6-AC-GFP transfer vector (OriGene) in-frame with a GFP tag at the C terminus or at the N terminus of the coding sequence. Viral particles were generated using the BacPAK baculovirus expression system (Clontech) and HEK 293 cells. 2–3 days after infection, Tip60 proteins were purified from the cells using a GST affinity column (Glutatione-agarose, bioprogen), eluted with 250 mM imidazole, and finally dialyzed against 20 mM Tris, 10% glycerol, and 1 mM dithiothreitol, at a pH of 7.5. The purified proteins were maintained at −20 °C and used in the PNGase F treatment experiment, mass spectrometry analysis, and the HAT assay.
Plasmids and Mutagenesis
The pCMV6-AC-GFP-Tip60 mutant strain (isoform 1; GenBank#Q5XI06) was purchased previously. The Tip60 mutant was generated via PCR and cloned in-frame with the GFP tag in the pCMV6-AC-GFP vector. The point mutations were generated by PCR, and the incorporation of all of the mutations was confirmed by DNA sequencing. In the Gln-342 (up 5'- att gat gga cgt aag Gac aag agt tat tcc -3', down 5'- ctg gga ata act ctt gtC ctt acg tcc atc -3'), Gln-324 (up 5'- cat cct cca ggc GCt gag att tac cgc aag -3', down 5'- gcg gta aat ctc aGC gcc tgg agg atg tcg-3'), and Gln-342/Gln-324 mutants the glutamine was replaced by alanine.
PNGase F Treatment
500 nanograms of GFP-Tip60 proteins produced in baculovirus were incubated with 10 units of PNGase F (PNGASE) (New England BioLabs) in the presence or absence of protease inhibitor (5 mM NaF) for 30 min at 37°C (or 1 h on ice when the deglycosylation was followed by a HAT assay). Tip60 was then removed via the incubation of the reaction mixture with NiTA-agarose beads and eluted with Laemmli sample buffer, then subsequently analyzed via SDS-PAGE and Western blotting. The PNGase F treatment of endogenous Tip60 was conducted as follows: HEK293 cell nuclei isolated from 107 cells were incubated for 30 min with 50,000 units of PNGase F at 37°C. The nuclei were washed three times in lysis buffer (15 mM NaCl, 60 mM KCl, 12% sucrose, 2 mM EDTA, 0.5 mM EGTA, 0.65 mM spermidine, 1 mM dithiothreitol, 0.5 mM phenylmethylsulfonyl fluoride, 0.5% Triton X-100) and lysed directly in protein loading buffer. Tip60 was detected with an anti-Tip60 antibody, as described by Legube et al.
Histone Acetyltransferase (HAT) ELISA Assays
Extracts were immunoprecipitated as above, except that the high salt wash was omitted. The immunoprecipitates were prepared test samples for each assay in 96-well plate. Mix the prepared assay mix, add 65µl of assay mix to each well, mix to start the reaction. Incubation plates at 37°C for 3hr depending on the color development. Read sample in a plate reader at 440nm (biovision). In some assays, HAT activity was analyzed with Tip60 immobilized on Concanavalin A beads after treatment with alkaline PNGase F.
Induction of ER Stress
ER stress was induced by exposure to brefeldin A. Brefeldin A is a nucleoside antibiotic that inhibits the N-glycosylation of target asparagine residues in the lumenal domains of proteins, whereas brefeldin A blocks the translocation of proteins from the ER to the Golgi apparatus by inducing the disassembly of the Golgi complex. Controls were treated with vehicle alone (dimethyl sulfoxide, 0.1%).
Cell Culture, Transfection, Drug Treatments, and Western Blotting
HEK293 cells were grown as previously described and transfected with EXGen500 (Fermentos) as indicated by the suppliers. 24-48 h later, the cell extracts were prepared by directly lysing cells in Laemmli sample buffer and sonication [20]. The transient Tip60-expressing HEK293 cells were cultivated in Dulbecco's modified Eagle's medium supplemented with fetal calf serum (10%) and antibiotics.
Confocal Microscopy
HEK293 cells were seeded overnight at 60% confluence onto culture slides coated with human fibronectin (Becton Dickinson, MA). The following day, cells were transfected TIP60 (WT) N324A, N342A mutants, and allowed to grow for an additional 48 hours. The cells were washed several times with ice-cold PBS and fixed in 2% paraformaldehyde for 10 minutes. The fixed cells were permeabilized with 0.1% Triton X-100 for 10 minutes and blocked for 2 hours in PBS containing 0.1% BSA-C (Aurion, The Netherlands) and 0.1% Tween. Following incubation with a polyclonal antibody against TIP60, the cells were washed and stained further with a conjugated donkey anti-rabbit IgG prior to processing the slides for immunofluorescence. After an additional 20 minutes of incubation at 37℃, the cells were fixed, permeabilized, and decorated with either an anti- TIP60 or GFP antibody. As a secondary antibody, Alexa Fluor 568-conjugated donkey anti-rabbit(Molecular Probes, Inc., Eugene, OR) was used. Confocal microscopy analysis was performed LSM710(Zeiss, Germany) at the Center for Experimental Research Facilities of Chungbuk National University.
Protein Stability Experiments
HEK293 cells (2.5 × 105 cells per well) in 10cm plates were transfected with 1.0 μg of expression with TIP60 WT and N324A mutant plasmid. The medium was replaced with medium containing 200 μg/ml cycloheximide 36 h after transfection (0-h time point). Cell lysates were harvested at 0, 8, 16, 24 h and analyzed by immunoprecipitation and Western blotting using anti-GFP antibody, and assayed in five time repeats. The relative optical density (OD) was measured by image analysis of the Fuji Image Quant software (Fujifilm, Tokyo, Japan), according to the manufacturer’s instructions.
RESULTS
The Association of Tip60 with Concanavalin A In Vivo and the Putative Site of Tip60 N-glycosylation
In order to determine whether Tip60 is an N- glycosylated protein, we treated HEK293 cell lysate human Tip60 with concanavalin A (Con A, Jack bean lectin) [21-23]. The extracts were then analyzed via Western blotting using an anti-Tip60 antibody (Fig. 1A left lane). Because Con A associates selectively with a-mannose-terminated glycans in solution and in its adsorbed state, we anticipated that a favorable interaction between Tip60 and Con A would be detected, if Tip60 was N-glycosylated in the HEK293 cells. To control the N-glycosylation of Tip60, we also added Con A to the HEK293 cell lysates, which had been pretreated with PNGase F [24]. We observed Tip60 in the Con A precipitant (Fig. 1A left lane), but not in the Con A precipitant after PNGase F treatment (Fig. 1A right lane). These findings indicated that Tip60 is one of the N-glycosylated proteins that react with Con A (a Jack bean lectin: Sigma).
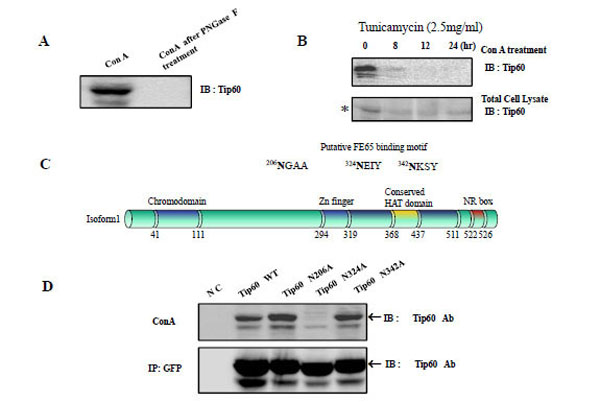
A) After treatment HEK293 cell lysate with concanavalin A (Con A, Jack bean lectin), the extracts were then analyzed by Western blotting using an anti -Tip60 antibody (A left). To control the N-glycosylation of Tip60, we also added Con A to HEK293 cell lysate was pretreated with PNGase F, before Con A treatment (A right). B) The reactivity of Tip60 to Con A was reduced with the tunicamycin (2.5mg/ml) treatment for different times, as indicated above (B upper lane). To control the amount of Tip60, the each cell lysate was immuno blotted with its specific antibody (B bottom lane). * indicated the putative glucosylated Tip60. C) With the http://www.cbs.dtu.dk/services/SignalP/output.html program, the putative N-glucosylation sites in Tip60 was predicted, as indicated in Fig. 1C above. The domains (chromodomain for the histone binding, Zn finger, the conserved HAT domain, NR for the nuclear receptor binding) are also indicated [6, 7]. D) After transfected with GFP Tip60 WT, N206A, N324A, N342A, the cell lysates was precipitated either Con A (D upper lane) or GFP Ab (D bottom lane). To detect Tip60 amount, immunoblot was performed with its specific Ab.
Tunicamycin is a nucleoside antibiotic that functions in eukaryotic cells to inhibit N-glycosylation by blocking the first step in the synthesis of the linked oligosaccharide: namely, the transfer of UDP-N-acetyl-glucosamine to dolichol-phosphate [25, 26]. We used tunicamycin to determine whether Tip60 glycosylation was affected by this drug. As is shown in Fig. 1B, the association of Tip 60 to Con A in HEK293 cells was attenuated with greater duration of tunicamycin treatment (2.5mg/ml); this suggests that Tip60 is modified via N-glycosylation. To monitor Tip60 expression amount by the tunicamycin treatment, the western blot with the Tip60 Ab was performed (Fig. 1B bottom). The putative N-glycosylated Tip60 (indicated as * marker in Fig. 1B bottom) form was reduced, depending on the tunicamycin treatment time.
We then attempted to identify the N-glycosylation site in Tip60. Using the http://www.cbs.dtu.dk/services/SignalP/output.html program (Fig. 1C), the putative N-glycosylated site was suggested on the Asn residues in (342NKSY345, where is well matched with N-glycosylation consensus sequences (Asn–X–Thr/Ser, where X is any amino acid except proline). Using site-directed mutagenesis we constructed Tip60 N206A or N342A, and attempted to determine whether these two were the putative N-glycosylated sites (Fig. 1D). We included Tip60 N206A, because this motif is matched with 173NGSA176 of Tip60 isform2 (NP_006379; Sterner and Berger. 2000; Squatrito et al. 2006). Also, we included the Tip60 N324A (in 324NEIY327) mutant where the motif is matched with the NxxY consensus motif. After GFP-tagged human Tip60 mutants were expressed in HEK 293 cells using the eukaryotic expression system, the blockage of N-glycosylation of the Tip60 N206A or N342A mutants failed (Fig. 1D), thus indicating that two Asn residues in (206NGAA209 and 342NKSY345) are not N-glycosylation sites. Rather, the Tip60 N324A (in 324NEIY327) mutant was not associated with Con A (Fig. 1D upper lane). To monitor Tip60 expression amount, the western blot with the Tip60 Ab was performed, after immunoprecipitating with GFP Ab (Fig. 1D bottom). Thus, unexpectedly, the 324 Asn residue appears to be the N-glycosylation site of Tip60 where does not match the consensus motif for the asparagine (N) residue of the NX(S/T).
Together with all these, Tip60 is one of the N-glycosylated proteins that react with Con A, and its 324 Asn residue in 324NEIY327 motif (where is located between the Zn finger and the conserved HAT domain) is the site for its N-glycosylation (Fig. 1C).
Con A Binding Site (A Putative Glycosylation Site) on Tip60 is Overlapped with the Interaction Site for FE65 Binding, and is Related with its HAT Activity
To characterize the effects of 324 Asn residue of Tip60 (in324NEIY327), we evaluated the protein-protein interaction occurring between the Tip60 mutant and FE65. The Fe65 PID domains are protein-protein interaction modules that recognize short peptide regions, including the canonical NPXpY consensus sequence (sometimes reduced to NXXpY; X indicates relaxed residue selectivity). PID domains differ in their specificities for phosphorylated or unphosphorylated tyrosines, and the majority of the analyzed PID domains bind their substrates in a Tyr(P)-independent manner. Previously, Cao and Südhof reported that histone acetyltransferase (Tip60) binds to Fe65 and forms a multimeric complex with Fe65 and the cytoplasmic tail of APP [27]. They also suggested that the 342NKSY345 motif of Tip60 binds with PID1 of Fe65.
In order to confirm their observation and elucidate the Tip60 glutamine 324 residue (Con A binding site), we immunoprecipitated the transfected GFP Tip60 WT, N324A, and N342A with GFP Ab, and applied with Tip60 or FE65 Ab for immune-blotting. As shown in Figs. (2A) and (B), Tip60 N324A only failed to co-immunoprecipitate FE65 in HEK293 cells, unexpectedly, suggesting that 324NEIY327 is the motif to bind with FE65 (Fig. 1C). This result also suggested that the 342NKSY345 motif in a loop of the histone acetyltransferase domain which was previously reported as the binding motif by Cao and Südhof is not responsible for binding to the PID 1 of FE65.
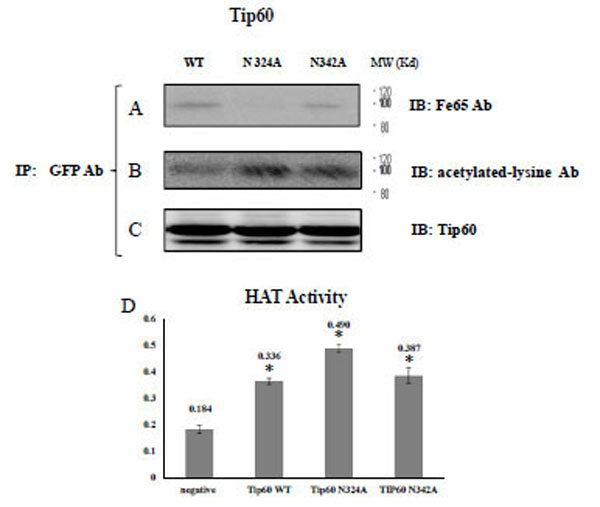
A) After immunoprecipitating the transfected GFP Tip60 WT, N324A, N342A with monoclonal GFP Ab, the precipitant was immuno blotted with FE65 Ab, B) Acetylated-lysine Ab, or C) Tip60 Ab. D) In parallel, HAT activity of purified Tip60 WT, N324A, or N342A expressed in HEK 293 cell were compared with ELISA method in arbitrary units. For the detail, see the material and method. * P < 0.05 significance versus to each other.
To monitor its own acetylation via self histone acetyltransferase activity, we conducted immune-blotting with an acetylated lysine-specific antibody (Fig. 2B). In parallel with the FE65 protein association shown in Fig. (2B), the acetylation of Tip60 N324A was well profound than was detected in the WT or N342A variants. Those observations suggest that the glycosylation site of Tip60 is involved in both its histone acetyltransferase activity and FE65 interaction (Fig. 2C). In an effort to evaluate further the role of glycosylation on 324 Asn on its histone acetyltransferase (HAT) activity, the HAT activity of purified Tip60 WT, N324A, or N342A expressed in HEK 293 cells was compared via ELISA (Fig. 2D). The results demonstrated that the HAT of Tip60 N324A was considerably reduced relative to the WT or N342 variants. This result suggests that glycosylation on the 324 Asn residue of Tip 60 is related with its HAT activity (Fig. 2D), and this is consistent with the results shown in Fig. (2C). Thus, a detectable change in HAT activity of Tip60 N342 was observed relative higher than that of WT control. The results of these experiments also demonstrate that the inhibition of Tip60 N324A HAT activity is not the result of mutation, but rather the consequence of its inability to be glycosylated. Our data revealed that the critical site of Tip60 glycosylation is also needed for the control of its catalytic activity (Fig. 2C and D). Collectively, the resutls of that analysis suggest that Tip60 HAT activity control seems to be related with the 324 Asn residue (Con A binding site), which also affects its binding to FE65. We also observed that FE65 expression level is reversible related with Tip60 HAT activity (data not shown). Further it appeared to be important to identify the glycosidase involved, in order to better understand the functional significance of this glycosylation-mediated regulation of Tip60 activity.
The Con A Binding Property (A Putative Glycosylation) of Tip 60 Enhances the Etoposide-induced G0-G1 Cell Cycle Arrest
To demonstrate the effect of Tip60 glycosylation on the cell cycle regulation, HEK293 cells were treated with Tip60 WT, N324A, or N342A, and the topo II poison etoposide. After selecting the transfected cells with GFP antibody, the cell cycle sorter was operated. Even though Tip60 WT has been reported to arrest G2-M phase [28, 29], we observed that this protein is effective on G0-G1 phase arrest. As shown in Fig. (3), either Tip60 N324A or N342A mutant appeared to be more effective to induce an arrest at the G0-G1 phase than Tip60 WT. Thus, these results suggested that the 324 Gln residue of Tip60 seems to be required for the enhancement of G0-G1 arrest of the cell cycle which was induced by the etoposide.
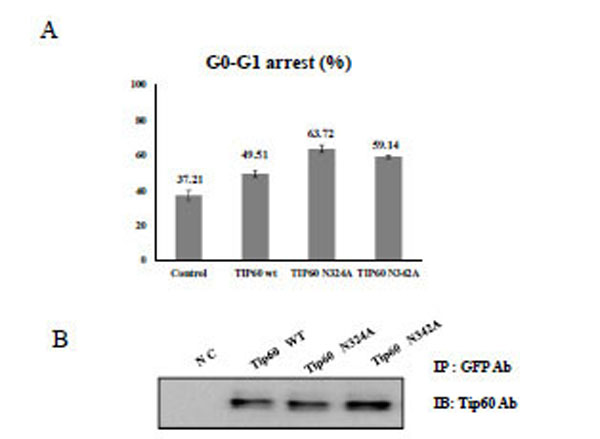
A) After transfection of HEK293 cells with each Tip60 construct as indicated below with Etoposide (5ug/ml), the G0-G1 cell cycle was sorted with the cell sorter. as a percent unit to the total cell numbers. The value represents the mean of five time repeats. B) Each Tip60 amount (indicated above) was monitored with its specific antibody.
The Protein Stability of Tip60 WT, or N324A
To compare the protein stability of Tip60 WT, or N324A, in HEK293 cells, Tip60 WT, or N324A, DNA were transfected in the cells. The transfected cells were treated with cyclohexamide and the protein stability was compared. The Tip60 proteins were chased for the indicated time periods. GFP Tip60 proteins were immunoprecipitated with a polyclonal anti-GFP antibody and subjected to SDS-PAGE followed by western blotting with a monoclonal Tip60 antibody (Fig. 4A). An equal amount of cell lysate was subjected to western blotting with an actin antibody to monitor the protein levels. As shown in Fig. (4B), Tip60 WT was more stable than N324A mutant. In order to eliminate the muta-tional effect of N324A mutant on its protein stability, the protein stability of Tip60 N342A mutant was also measured with the same procedure. The protein stability of Tip60 N342A mutant was almost same as that of Tip60 WT (data not shown). Thus, these data also indicated that 324 Gln residue of Tip60 seems to be required for its protein stability.

A) GFP-Tip60 or Tip60 N324A mutant was transfected into HEK293 cells and the cells were treated with cyclohexamide. The Tip60 proteins were chased for the indicated time periods. Western blotting with a monoclonal Tip60 antibody of SDS-PAGE subjected GFP-Tip60 proteins is shown. To monitor the protein amount, an equal amount of cell lysate was subjected to western blotting with an actin antibody. Results shown are one of five repeated experiments. B) Quantification of the pulse-chase experiment is shown by image analysis with the Fuji Image Quant software.
The Glutamine 324 Residue Mutation of Tip60 Affects on Its Subcellular Localization and Co-localization with FE65
In an attempt to determine whether or not the observed change in the Gln 324 residue of Tip60 is attributable to its subcellular localization or to deviation of FE65 co-localization, we performed confocal microscopic analysis with transfected GFP-Tip60 WT and N324A, N342A (all green color) to the endogenous FE65 (all red color). Each was compared with one another via direct immunofluorescence microscopy (Fig. 5). Tip60 N324A was detected predominantly in the nuclear (middle lane), whereas the Tip60 WT or N324A was localized both cytoplasm and nuclear (upper and bottom lane). The significant difference in the subcellular localization of Tip60 N324A from that of Tip60 WT or N342A seems to be related with its Con A binding (Fig. 1D), suggesting that this modification of Tip60 promotes its ER retention.
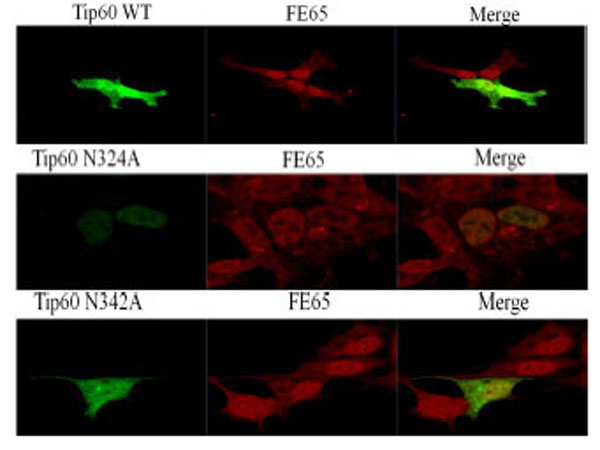
The confocal microscopic images transfected with GFP-Tip60 WT, N324A N342A (all green color) to the endogenous FE65 (all red color) were compared each other in the direct immunofluorescence microscopy. Results shown are one of five repeated experiments.
To further confirm that Gln 324 residue of Tip60 is required to bind with FE65 (Fig. 2A), we attempted to determine whether Tip60 N324A (middle lane) is disrupted to its co-localization with the endogenous FE65 (red color). As is shown in Fig. (5), both Tip60 WT (upper lane) and N342A (bottom lane) were merged well FE65 in the nuclear (yellow color). Meanwhile, we noted a significant difference of the co-localization of Tip60 N324A with FE65 (middle lane). Even though Tip60 N324A was found in the nuclear, it was not merged well with FE65 in the nuclear, comparing with Tip60 WT and N342A. Thus, Gln 324 residue mutation of Tip60 which ablates its Con A binding (a putative N-glycosylation) seems to promote positively its nuclear localization with reducing its ER retention, while this mutant disturbs the co-localization with FE65.
Brefeldin which Induces ER Stress, Increases Con A Binding of Tip60
In an effort to characterize the biological functions of Tip 60 Con A (lecithin) binding, we hypothesize that the property is associated with the ER stress signal, because N-glycosylation occurs in the ER or Golgi apparatus. Because it has been previously noted that brefeldin or thapsigargin A induce ER stress, we used brefeldin to induce ER stress in HEK293 cells [30-32].
Consequently, with the administration of brefeldin to HEK293 cells, Tip60 Con A binding was increased in a time-dependent or concentration-dependent manner (Fig. 6 upper lane). The induction time to the saturated Con A binding was shorten from 24hr at 2.5 mg/ml of brefeldin (Fig. 6 left lane) to 8hr at its 10 mg/ml (Fig. 6 right lane) The Con A binding of Tip60, which was detected by Con A bead binding, increased depending on both its treatment time and concentration (Fig. 6 upper lane), without increasing Tip 60 total amount (Fig. 6 bottom lane). The brefeldin (an ER stress inducer) seemed to enhance Tip60 to Con A beads through its N-glycosylation. We also observed the increase of Tip60 to Con A binding with the thapsigargin A treatment (data not shown). Therefore, even though the precise relationship between the glycosylation of Tip60 and ER stress remains unknown, the Con A binding of Tip60 (or glycosylation on its 324 Gln residue) seems to be related with one of signaling steps for the ER stress.
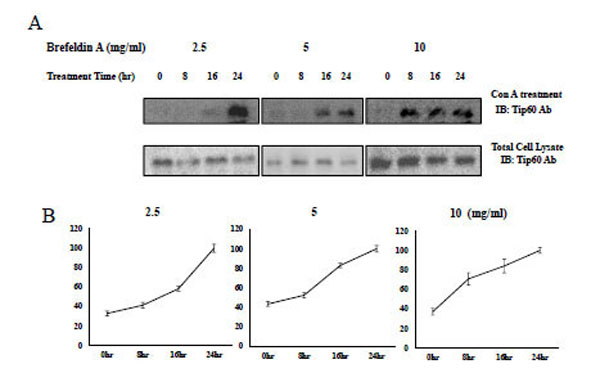
A) After treatment of brefeldin to HEK293 cell in the indicated time and concentration, Tip60 glycosylation, the glycosylation of Tip60 which was detected by Con A bead (upper lane). For the control, each sample was immunoblotted with Tip60Ab (bottom lane). Result shown is one of five repeated experiments. B) The relative quantification of Tip60 glycosylation increase with brefeldin treatment is shown by image analysis with the Fuji Image Quant software. The relative arbitrary unit to the maximum value on the specific time in each brefeldin treatment concentration (indicated above) was shown.
In summary, we demonstrate in this report that Tip 60 is a Con A binding protein (through the putative N-glycosylation on its 324 Gln residue), which seems to influence on its stability, protein interaction, enzyme activity, cell cycle regulation, and subcellular localization (or ER retention). Thus, the Con A binding of Tip60 which is responsible to ER stress appears to regulate its several functions.
DISCUSSION
The principal objective of this paper was to ascertain whether Tip60 is glycosylated and to evaluate the role of Con A binding (a putative N-glycosylation) site in Tip 60 HAT activity. Consequently, using site-directed mutagenesis we have identified the Con A binding (N-glycosylation ) site as the Asn 324 residue, which is normally occupied during Tip 60 HAT activity, and it is required for binding FE65 in HEK 293 cells (Figs 1-6). We measured the enzymatic activity of wild-type and recombinant Tip 60 mutant HAT activity and evaluated the kinetics of folding and Con A interaction. These results show that the Con A binding ability of Tip 60 play distinct roles in the Tip 60 HAT activity folding kinetics, suggesting a local folding mechanism for this glycoprotein. Here, we demonstrate that the lecithin binding of Tip60 is a controlled phenomenon, and modulates the activity of the protein (Fig 1-6).
In N-linked glycosylation, a carbohydrate is linked covalently to an asparagine residue within the consensus sequence Asn–X–Thr/Ser, where X is any amino acid except for proline [12, 13]. The putative N-glycosylated sites are suggested on two Asn residues in (206NGAA209 or 342NKSY345, where is well matched with N-glycosylation consensus sequences (Asn–X–Thr/Ser, where X is any amino acid except proline). Unexpectedly, with site directed mutagenesis, we demonstrated here that Tip60 324 Asn (in 324NEIY327) is responsible for its N-glycosylation (Fig. 1). However, we do not exclude that either 206NGAA209 or 342NKSY345 is also N-glycosylated sites during its translation process in ER.
N-linked glycosylation occurs co-translationally, and the addition of the carbohydrate to the partially folded, nascent polypeptide can impact, or facilitate, the process of protein-folding [11, 14]. Thus, it is possible that N-glycosylation on 324 Gln of Tip60 is what is left of its translation process in ER. It is also interesting to note that even though Tip60 is known as a nuclear protein but not a membrane associated protein, this protein is present usually as a glycosylated form in the cell which can easily bind to Con A lectin. However, Tip60 glycoprofile remains to be solved, and the origin and process of N-glycosylation on its 324 Gln residue is not known at present time.
In our experiments, we observed that the glycosylation of Tip60, which was detected by Con A bead binding, increased depending on treatment time and concentration of brefeldin (Fig. 6A), even though the precise relationship between the glycosylation of Tip60 and ER stress remains to be characterized [14-16]. Because the chemical (such as brefeldin) induced the glycosylation of Tip60 without increasing Tip60 expression, ER stress seems to increase Tip60 glycosylation for the binding with ER lectin, involving in the calnexin/ calrecticulin cycle to link ERAD substrate, or blocking the retro-translocation from ER lumen to the cytoplasm [14-16]. We will pursue further to define this possibility that ER stress increases Tip 60 glycosylation for its ER retention, affecting on its stability, HAT activity, FE65 interaction, and G0-G1 arrest (Figs. 1-4). The precise relationship between the glycosylation of Tip60 and ER stress remains to be characterized whether the glycosylation of Tip60 is a result of ER stress, or is a step of ER stress signaling. It also remains unclear as to whether such ER stress increases Tip60 glycosylation concurrently with the ER stress signal pathway, as demonstrated above. In present time, we hypothesize that ER stress enhances the glycosylation of Tip60 in order to promote the apoptosis or the cell cycle arrest through reducing its nuclear localization.
FE65 is capable of interacting with the β-amyloid precursor protein (APP) through the PID2 of FE65, as well as with other members of the APP protein family. The PID2 domain of FE65 binds the YENPTY sequence in the cytoplasmic domain of APP [27, 33, 34]. Because FE65 operates as an adaptor protein within the cell, the identification of ligand proteins for the PID1/PID2 domain or the WW domain may provide us with some clues to its biological functions. Further, it may also be important to establish the manner in which the protein-protein interaction between FE65 and its ligand protein (such as Tip60) is regulated by posttranslational modifications, such as protein glycosylation, acetylation, or phosphorylation [35, 36]. Because of the overlapping Tip60 glycosylation site and its FE65 binding motif, the protein interaction between Tip60 and FE65 seems to be affected by Tip60 glycosylation (Fig. 2A). Further, FE65 seems to inhibit Tip60 HAT activity through binding its PID1 domain with 324NEIY327 motif of Tip60, where is overlapped with Tip60 glycosylation site. We are now perusing the possibility that FE65 negatively regulates HAT activity of Tip60 (Fig. 2C and D). Recently, other researchers reported that Tip60 is regulated through auto-acetylation. Similarly, it seems to be that the dissociation Tip60 from FE65-Tip60 protein complex activated its HAT activity, as observed with Tip60 N324A.
In conclusion, this study established a link between the Con A binding property of Tip60 (which seems to be regulated by N-glycosylation on 324 Gln residue with ER stress) and its ER retention, stability, HAT activity, or interactions with FE65. However, even though the glycosidase responsible for the glycosylation of Tip60 remains elusive, the elucidation of its specific posttranslational modification (glycosylation or deglycosylation) with one or several of the signaling pathways in a step-by-step manner will help us to understand its biological functions and the mechanism underlying its regulation.
CONFLICT OF INTEREST
None declared.
ACKNOWLEDGEMENTS
This work was supported by National Research Foundation of Korea (NRF) grants (2009-0076024 and 2009-0069007) funded by the Korea government (MEST) to S.S. Kang. We also appreciate the help of The Core Facility of Chungbuk National University.
REFERENCES
[PubMed Link]
[PubMed Link]
[PubMed Link]
[PubMed Link]
[PubMed Link] [PMC Link]
[PubMed Link]
[PubMed Link] [PMC Link]
[PubMed Link]
[PubMed Link] [PMC Link]
[PubMed Link] [PMC Link]
[PubMed Link]
[PubMed Link] [PMC Link]
[PubMed Link] [PMC Link]
[PubMed Link]
[PubMed Link] [PMC Link]
[PubMed Link]
[PubMed Link]
[PubMed Link]
[PubMed Link] [PMC Link]
[PubMed Link]
[PubMed Link]
[PubMed Link]
[PubMed Link]
[PubMed Link]
[PubMed Link]
[PubMed Link]
[PubMed Link]
[PubMed Link]
[PubMed Link]
[PubMed Link] [PMC Link]


