All published articles of this journal are available on ScienceDirect.
Vitamin D and Multiple Sclerosis: An Open-Ended Story
Abstract
Multiple Sclerosis (MS) is a chronic inflammatory autoimmune disease of the Central Nervous System (CNS). Genetic, epigenetic and environmental factors interact together, contributing to the complex pathogenesis of the disease. In the last decades, the role of hypovitaminosis D on MS risk was hypothesised. Several factors drive the regulation of vitamin D status, including genetics. The current review summarises the literature evidence on the association between vitamin D and MS, with a focus on the genetic polymorphisms in vitamin D-related genes. The variants of the genes codifying Vitamin D Receptor (VDR), Vitamin D Binding Protein (VDBP) and CYP enzymes have been investigated, but the findings are controversial. Only a few studies have addressed the role of DHCR7 polymorphisms in MS risk.
1. INTRODUCTION
Multiple Sclerosis (MS) is an autoimmune disease of the Central Nervous System (CNS). The triad consisting of inflammation, demyelination and neuroaxonal damage in the CNS causes clinical disability, characterised by a wide spectrum of signs and symptoms. MS is a multifactorial disease, the pathogenesis of which is the result of the interaction among genetic, epigenetic and environmental factors [1].
In the last decades, hypovitaminosis D has emerged as a risk factor for MS. Initially, since vitamin D is mainly produced in the skin by the action of Ultraviolet (UV) radiation [2], it was proposed as the scapegoat to explain the association between reduced solar radiation exposure and increased risk of developing MS. Indeed, an inverse correlation between geographical latitude and MS prevalence, with increasing risk from the equator to North and South Poles, has been reported [3]. However, this hypothesis is still debated. Koch-Henriksen and Sørensen reported a latitudinal gradient of incidence of MS in Europe and North America, but not in Australia and New Zealand [4]. Also, Bezzini and Battaglia observed an inverse correlation between MS and latitude in Europe [5]. However, they concluded that the increasing prevalence could be the result of increasing survival and incidence. Overall, many profound cultural, economic, and, sociological differences between northern and southern Europe can influence the geographical distribution of MS; for instance, viral and fungal infections may be related to MS spreading among selected individuals, including drug users and homosexuals/bisexuals [6, 7]. In particular, Benito-Leon and Laurence suggested that MS could be due to a fungal pathogen because vitamin D related gene variants are involved in fungal diseases [6].
Collective data from epidemiological, clinical and experimental studies suggest that genetics also contribute to the vitamin D status, with an estimated heritability ranging from 23 to 80% [8]. In particular, genetic studies disclosed several Single Nucleotide Polymorphisms (SNPs) in the genes codifying molecules involved in the vitamin D pathway associated with the vitamin D status. Accordingly, several authors investigated the influence of these SNPs on MS risk.
The current review summarises the literature evidence on the association between vitamin D status and MS risk.
2. VITAMIN D
2.1. Metabolism
Vitamin D, either in D2 or D3 form, is a secosteroid prohormone. Vitamin D3 is mainly produced endogenously in the skin where Ultraviolet B (UVB) rays convert 7Dehydrocholesterol (7DHC) into pre-vitamin D3 that, spontaneously, isomerizes into vitamin D3. However, it can be also introduced exogenously [9-11]. Vitamin D2 is produced by UVB radiation of the ergosterol in fungi; it can be introduced in humans by the diet, including food, supplementation and fortification [12]. Vitamin D3, both endogenous and exogenous, as well as vitamin D2, is a biologically inactive prohormone that becomes an active hormone after two hydroxylation reactions. The first 25-hydroxylation reaction takes place in the liver and produces 25-hydroxyl-vitamin-D2 [25(OH)D2] and 25-hydroxyl-vitamin -D3 [25(OH)D3]. The second hydroxylation reaction converts 25(OH)D2 and D3 in the biologically active form, 1,25-dihydroxy-vitamin-D2 [1,25(OH)D2] and 1,25-dihydroxy-vitamin-D3 [1,25(OH)D3]; this reaction occurs mainly in the kidney proximal tubule cells. From now, the terms 25(OH)D and 1,25(OH)2D will be used to indicate the sum of D2 and D3 metabolites. Vitamin D binding protein (VDBP) binds both 25(OH)D and 1,25(OH)2D with high affinity [13].
1,25(OH)2D exerts both genomic and non-genomic actions through the interaction with vitamin D receptor (VDR) and Membrane-Associated Rapid Response Steroid-Binding proteins (MARRS), respectively [14]. Besides its role in calcium metabolism, it is now widely known that vitamin D exerts several pleiotropic effects regulating different biological functions, including the immune response.
Vitamin D catabolism depends on carbons C-24 and C-23 hydroxylation reactions.
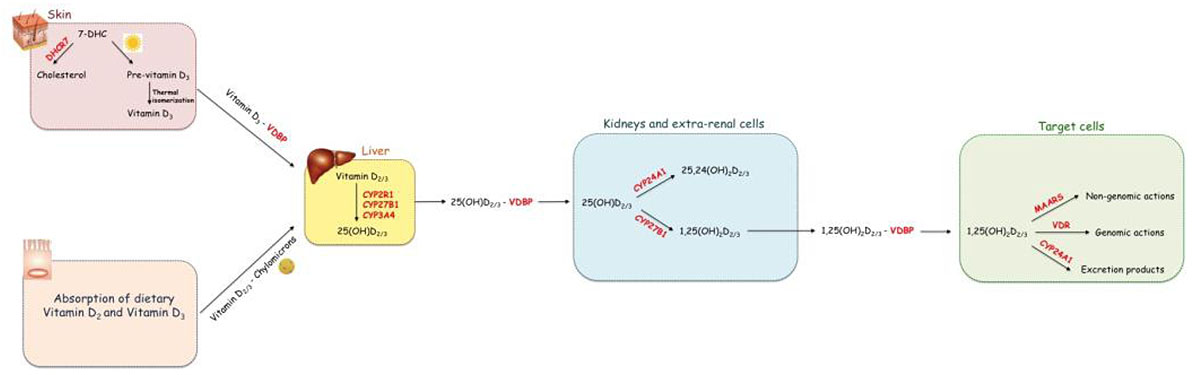
The three main steps in vitamin D metabolism, namely 25-hydroxylation, 1α-hydroxylation, and 24-hydroxylation, are all executed by cytochrome P450 enzymes (CYPs) [14, 15]. (Fig. 1) shows the metabolic pathway of vitamin D.
Vitamin D, both D2 and D3, can be either synthetized in the skin or absorbed by enterocytes forming cholecalciferol. It becomes active Vitamin D after two hydroxylations steps. The first hydroxylation occurs in the liver producing 25(OH)D; the second hydroxylation takes place in the kidney and in extra-renal cells and produces the biologically active form of vitamin D, 1,25(OH)2D. The latter exerts its biological actions through the interaction with the intracellular receptor VDR or the transmembrane receptor MARRS.
2.2. Vitamin D Status and its Heritability
Vitamin D status is assessed in clinical practice by the measurement of serum 25(OH)D levels, which are considered the best indicator of circulating vitamin D. The reasons for this can be summarised as follows: i) 25(OH)D has a longer half-life than 1,25(OH)2D (2–3 weeks vs 4-6 hours); ii) while 25(OH)D levels are unregulated, 1,25(OH)2D levels are tightly regulated and strictly kept between limits even when adverse effects start to occur; iii) 1,25(OH)2D can be produced both in the kidney cells and in many other cells. The extra-renal 1,25(OH)2D does not enter into the circulation, but it acts locally. Thus, serum 1,25(OH)2D does not reflect the total 1,25(OH)2D. In contrast, circulating 25(OH)D mirrors 25(OH)D reserve prior to the conversion, also including the available substrate for the extra-renal 1,25(OH)2D synthesis [16].
Extensive research over the past decades has suggested that genetics greatly influences vitamin D status [4], as reported by twin and family studies. To date, numerous variants in genes codifying molecules involved in the vitamin D metabolism and associated with altered circulating vitamin D levels have been identified. This evidence is derived from genetic studies based mainly on two different approaches: candidate gene study and Genome Wide Association Study (GWAS) [4]. The candidate gene approach is based on the selection of a gene depending on the knowledge of the metabolic pathway or pathogenesis disease. GWAS, instead, is a hypothesis-free approach that evaluates the entire genome to identify allelic variants that are associated with a specific trait [17, 18]. Both the approaches have advantages and disadvantages that have been well summarised in the review by Lin [19]. However, both are pivotal in understanding the genetics of vitamin D status.
3. MULTIPLE SCLEROSIS
MS represents a common neurological disease in young adultswhich is usually diagnosed between the third and fifth decade of life. It is characterized by sexual dimorphism influencing both the incidence and severity of the disease. In particular, the prevalence is higher in women than in men, with a female to male ratio of 2:1-3:1, and a “maternal parent-of-origin” effect. However, men present an overall worse course of the disease, with a rapid accrual of disability and progression [20].
MS is an immune-mediated disease characterized by the gradual destruction of myelin due to an autoimmune response against self-antigens [21]. The characteristic pathological hallmark of MS is the inflammation in the CNS, leading to demyelinating plaques localized mainly in the periventricular area, pons and spinal cord [22]. MS has long been considered a T cell-mediated inflammatory disease [23, 24]. However, the detection of the Oligoclonal Band (OCB) in the CSF of MS patients and the success of B-cell targeted therapies challenge the standard T-cell autoimmune dogma [25]. Studies on Experimental Autoimmune Encephalomyelitis (EAE), which represents the best animal model of MS, have made a great contribution to understanding the pathogenesis of MS. It is now well documented that both CD4 and CD8 T lymphocytes and their related cytokines, as well as B-lymphocytes, take part in the development of MS. Among CD4 T cells, activated myelin-specific T helper (Th)1 and Th17 with their related cytokines, such as interferon-gamma (IFN-γ), interleukin (IL)-17, IL-22 and granulocite-macrophage colony-stimulating factor (GM-CSF), play a pivotal role in initiating the inflammation in the CNS [26, 27]. B-lymphocytes contribute to the MS pathogenesis through different mechanisms including the production of antibody-producing plasma cells, potent Antigen-Presenting Cells (APC) and secretion of pro-inflammatory cytokines [25]. Moreover, chronic stages of MS are characterized by a compartmentalized immune response in the CNS with activated microglia and macrophages [28].
Although MS is not a hereditary disease, genetic predisposition increases the risk of developing it [29-31]. A meta-analysis of twins revealed that genetic alterations could at least partly explain individual differences in MS susceptibility [32]. Epidemiological studies suggested that the individual risk for subjects with a family member affected increases proportionally to the degree of kinship and, consequently, to the percentage of shared genetic background, being the highest among monozygotic twins (25-30%) and greatly decreasing in siblings (5%), parents (2%) and distant relatives (1%) [33]. The Common-Disease-Common Variant (CD-CV) hypothesis seems to be the best model to explain MS heritability [34]. According to this model, MS risk is the consequence of the cumulating effects of numerous allelic variants, which are common in the general population.
Genetic studies revealed variants in several genes associated with MS, including those codifying molecules involved in vitamin D metabolism. CYP27B1 and CYP24A1 were the first genes involved in vitamin D metabolism identified by a GWAS, including 9772 MS and 17376 controls [35]. This finding further supported the importance of vitamin D in MS risk and represented the foundation for investigating the influence of variants in genes implicated in vitamin D metabolism, catabolism and function on MS.
4. VITAMIN D AND MULTIPLE SCLEROSIS
4.1. Pathogenic Role of Hypovitaminosis D in Multiple Sclerosis
Vitamin D has been long investigated in many inflammatory- based, neurological, autoimmune and infectious diseases either as a risk factor or biomarker of severity, along with other biomarkers [36-40]. In particular, 25(OH)D has been recently proposed as a serum biomarker in neurodegenerative disorders, together with cardiovascular and inflammatory markers [41-51].
Compelling evidence from epidemiological studies, experimental studies on animal models, in vitro, observational clinical and genetic studies suggest a role of hypovitaminosis D on MS risk and progression [52-59]. This evidence is supported by the immunoregulatory and neuroprotective function of vitamin D [60].
Significantly, immune cells, including macrophages, T and B lymphocytes, express the enzymatic machinery required to metabolize vitamin D, namely CYP27B1 and CYP24A1, and the receptors making them sensitive to its action. In particular, vitamin D regulates the immune response through several mechanisms that can be summarised as follows: i) inhibition of the synthesis and secretion of pro-inflammatory cytokines; ii) inhibition of Th1 and Th17 cell differentiation; iii) promotion of regulatory T-cells (Tregs) and macrophages proliferation; iv) inhibition of proliferation and differentiation of plasma cells as well as antibody production by B cells. Consequently, hypovitaminosis D shifts the balance of the immune system toward a pro-inflammatory state [61, 62]. Further details on the relationship among vitamin D, the immune system and MS are provided in a recent review by Lu et al. [63].
Vitamin D is also regarded as a neurosteroid [64]. Studies on CSF revealed that both 25(OH)D and VDBP are detectable within the CNS and most CNS cells, including neurons, astrocytes and microglia cells, express both VDR and CYP27B1 [65-69]. Thus, all this evidence supports a direct action of vitamin D on some functions of the CNS. Some authors described the role of vitamin D in the myelination and remyelination processes [70-74]. As previously mentioned, MS is characterized by progressive demyelination. However, in the early stage of MS, demyelinated lesions undergo a partial remyelination process as shown by neuroimaging findings [71]. Some authors showed that vitamin D is involved in the demyelination/ remyelination process, by modulating the differentiation of the Oligodendrocyte Precursor Cells (OPC) [70]. Moreover, vitamin D stimulates microglia activation, promoting the clearance of myelin debris and favouring remyelination [72].
The inverse association between vitamin D and MS risk is supported by numerous observational studies conducted both on children and adults worldwide, from Australia to Europe [75-78]. The causal relationship between hypovitaminosis D and MS susceptibility is also proved by Mendelian Randomization (MR) analysis [79]. Usually, Randomized Controlled Trials (RCTs) represent the gold standard for exploring causal inference between certain exposures and disease. However, an MS prevention trial with vitamin D is impossible for several reasons, such as the low incidence of the disease in the population and a long follow-up starting from the prenatal period to the fourth decade of life [80]. Alternately, MR represents a powerful tool to verify the causal association between a certain trait, such as vitamin D, and an outcome, such as Multiple Sclerosis, based on genetic associations. Recently, three studies based on this epidemiological approach provided strong evidence of the causal role of low vitamin D levels in MS, even after accounting for other risk factors, such as smoking, and other potential confounders, such as sun exposure and diet [59, 79, 81].
Another evidence supporting the pathogenic role of hypovitaminosis D in MS is that vitamin D could regulate most of the MS-associated genes through the interaction with the Vitamin D Responsive Elements (VDREs) located within their promoter region [82, 83].
Collectively, all these data support the role of hypovitaminosis D in MS. This lead to the investigation of the effect of vitamin D supplementation on MS. However, the findings have not been encouraging as shown by some authors [84-86]. A recent meta-analysis reported inconclusive results showing no effect of vitamin D on clinical outcomes in MS patients [84]. Other authors reported the effect of vitamin D supplementation on improving mental health in MS patients finding questionable results [85]. Also, the effect of vitamin D supplementation on inflammatory markers has been studied in MS patients, resulting in no difference between vitamin D and the placebo group [86]. Thus, further studies are needed to clarify whether vitamin D supplementation could be useful in MS patients.
4.2. Variants in Vitamin D Related-genes and Multiple Sclerosis
Polymorphisms in most of the genes involved in the vitamin D pathway have been investigated in MS. It should be noted that vitamin D related gene variants are associated with many biological phenomena, other than MS onset [6].
7-dehydrocholesterol reductase (DHCR7) gene variants were first associated with hypovitaminosis D in 2010 by two independent GWAS [87, 88]. DHCR7 gene encodes an enzyme that catalyses the last reaction in the cholesterol biosynthesis pathway using 7-DHC as a substrate. 7-DHC is a common precursor of vitamin D3 and cholesterol. Indeed, in the skin, 7-DHC can be converted into vitamin D3 by UVB radiations. The conversion of 7-DHC by DHCR7 removes the substrate (7-DHC) from the synthetic pathway of vitamin D3. The results from GWAS identified genetic variants in the DHCR-7 gene as strong determinants of circulating 25(OH)D levels [87, 88] and several evidence support this association [89-91]. However, only a few studies addressed the influence of allelic variation on this locus and MS [85-88], with contrasting results. The influence of DHCR7 on MS risk requires further investigations.
The first hepatic hydroxylation of vitamin D can be catalysed by CYP2R1, CYP27A1 and CYP3A4. Among these enzymes, CYP2R1 has a prominent role in vitamin D metabolism, while CYP27A1 and CYP3A4 are also involved in the hydroxylation of other substrates.
The association between CYP2R1 and hypovitaminosis D was first identified by GWAS [87, 88]. Based on this evidence, case-control studies were performed to investigate a possible link between CYP2R1 polymorphisms and MS risk [92-97], with encouraging results.
The second hydroxylation of vitamin D is catalysed by CYP27B1 in many cells and tissues [98].
The association between CYP27B1 and MS risk was first identified by a GWAS performed by the Australia and New Zealand Multiple Sclerosis Genetics (ANZgene) Consortium [99]. Such association has been confirmed by some observational case-control and family studies [35, 93, 96, 100-105].
The levels of both 25(OH)D and 1,25(OH)2D are tightly regulated by CYP24A1, which mediates a hydroxylation reaction at C-24 and C-23, producing inactive metabolites.
The transcriptional expression of CYP24A1 is regulated by 1,25(OH)2D through a negative feedback mechanism. Indeed, vitamin D induces mRNA expression by the interaction with two VDREs in the promoter region of CYP24A1.
Simon et al. first explored the association between SNPs in the CYP24A1 gene and MS risk in a nested case-control study, achieving inconsistent results. The role of CYP24A1 on MS genetic susceptibility was first detected by a GWAS [35]. However, subsequent studies sometimes fail to replicate this finding [94, 102, 106-108]. Hence, the role of CYP24A1 polymorphisms in MS is still controversial.
Another protein with a key role in the vitamin D pathway is the VDBP, also known as Group-specific component (Gc), which regulates the vitamin D availability to target cells [109]. It belongs to the albumin gene family and it is synthetized in the liver and secreted in the blood where it binds most of the circulating vitamin D metabolites with high affinity (85-90%). VDBP also promotes the access of vitamin D to target cells through the interaction with transmembrane receptors, namely megalin and cubilin, which internalize the VDBP-vitamin D complex. VDBP also plays several other biological functions including scavenging action, fatty acid transport and macrophage activation [110].
The GC gene, which codifies VDBP, is localized in chromosome 4 and extends over 35kb; it consists of 13 exons and 12 introns. GC is one of the most polymorphic genes of the whole genome [109]. Among all the polymorphisms identified in the GC gene, two non-synonymous SNPs in the exon 11, the rs7041 and the rs4588, have been widely investigated in MS due to their influence on the vitamin D status, achieving conflicting results [93-95, 106, 111-115].
As above-mentioned, the genomic actions of vitamin D are mediated by VDR. It is a member of the nuclear receptor family of ligand-activated transcription factors, mainly localized in the cytoplasm. After binding 1,25(OH)2D, cytosolic VDR interacts with retinoid-X-receptor α (RXRα). The 1,25(OH)2D-VDR-RXRα complex moves into the nucleus where it regulates the expression of several genes through the interaction with VDRE in the promoter region of target genes [63, 116]. The following SNPs in the VDR gene have been widely investigated in MS: ApaI (rs7975232), BsmI (rs1544410), TaqI (rs731236), and FokI (rs2228570), achieving contrasting and contradictory results [96, 117-141]. While ApaI, BsmI and TaqI influence the VDR gene expression, FokI is associated with alterations of both protein structure and transcriptional activity of VDR (for a recent review of the association between VDR SNPs and the risk of MS see ref 142). While Tizaoui et al. reported an association of VDR polymorphisms with an increased risk of MS in large meta-analyses [143], Huang et al. [144] and Zhang et al. [142] conducted two meta-analyses showing opposite results. The reasons for this discrepancy could be the small sample size of most studies and, consequently, low statistical power, interactions with other genetic or environmental factors as well as clinical heterogeneity. Overall, the role of VDR genetic variations in MS risk must still be elucidated. The influence of RXR-α SNPs on MS risk has been evaluated in very few studies that found no association [95, 107]
MARRS, the receptor that mediates the non-genomic actions of vitamin D, is expressed in caveolae-enriched plasma membranes of target cells [145, 146]. To date, only one study investigated the influence of SNPs in the gene codifying for MARRS on MS risk, achieving inconsistent results [95].
CONCLUSION
The role of hypovitaminosis D in MS risk has emerged in the last decades and is supported by robust literature evidence. Notably, genetic background influences vitamin D status. Thus, several authors investigated the relationship among MS, vitamin D status and SNPs in genes involved in the vitamin D pathway. However, contradictory findings have been observed. This could be due to study limitations, including heterogeneity in the study design and small sample size. Moreover, standardization of 25(OH)D measurement is still lacking, which hampers the development of consensus guidelines about vitamin D deficiency, insufficiency and sufficiency. Thus, literature data does not allow drawing conclusions on the association among vitamin D status, genetics and MS. Further studies are required to better understand the role of vitamin D- related gene variants in the risk and progression of MS.
LIST OF ABBREVIATIONS
| CNS | = Central Nervous System; |
| UV | = Ultra Violet ; |
| SNPs | = Single Nucleotide Polymorphisms; |
| UVB | = Ultraviolet B; |
| 25(OH)D | = 25-hydroxyl-vitamin-D; |
| 1,25(OH)D | = 1,25-dihydroxy-vitamin-D; |
| VDR | = Vitamin D Receptor; MARRS, Rapid Response Steroid Binding Proteins; |
| CYPs | = Cytochrome P450 Enzymes; |
| GWAS | = Genome Wide Association Study; |
| OCB | = Oligoclonal Band; |
| EAE | = Experimental Autoimmune Encephal Omyelitis; |
| Th | = T Helper; |
| IFN-γ | = Interferon Gamma; |
| IL | = Interleukin; |
| GM-CSF | = Granulocite-Macrophage Colony-Stimulating Factor; |
| APC | = Antigen Presenting Cells; |
| CD-CV | = Common-Disease-Common Variant; T-regs, Regulatory T-cells; |
| OPC | = Oligodendrocyte Precursor Cells; |
| RCT | = Randomized Controlled Trials; |
| MR | = Mendelian Randomization; |
| VDREs | = Vitamin D Responsive Elements; |
| DHCR7 | = 7-Dehydrocholesterol Reductase; |
| GC | = Group-Specific Component; |
| 7-DHC | = 7-Dehydro-Cholesterol; |
| 25,24(OH)2D2/3 | = 25,24-Dihydroxy-Vitamin-D2/3; |
| 1,25(OH)D-26,23lactione | = 1,25-Dihydroxy-Vitamin D-26,23lactione; |
| MARRS | = Membrane-Associated Rapid Response Steroid Binding Proteins. |
CONSENT FOR PUBLICATION
Not applicable.
FUNDING
None.
CONFLICT OF INTEREST
The authors declare no conflict of interest, financial or otherwise.
ACKNOWLEDGEMENTS
Declared none.


